Congratulation to our friend and colleague Melanie Föll from the University of Freiburg Oliver Schilling lab to her great article on the analysis of mass spectrometry imaging data. She has integrated 18 tools dedicated to mass spectrometry imaging into our Galaxy framework. Thanks a lot for making your tools available to the proteomics and imaging community!
Abstract - Mass spectrometry imaging is increasingly used in biological and translational research because it has the ability to determine the spatial distribution of hundreds of analytes in a sample. Being at the interface of proteomics/metabolomics and imaging, the acquired datasets are large and complex and often analyzed with proprietary software or in-house scripts, which hinders reproducibility. Open source software solutions that enable reproducible data analysis often require programming skills and are therefore not accessible to many mass spectrometry imaging (MSI) researchers.
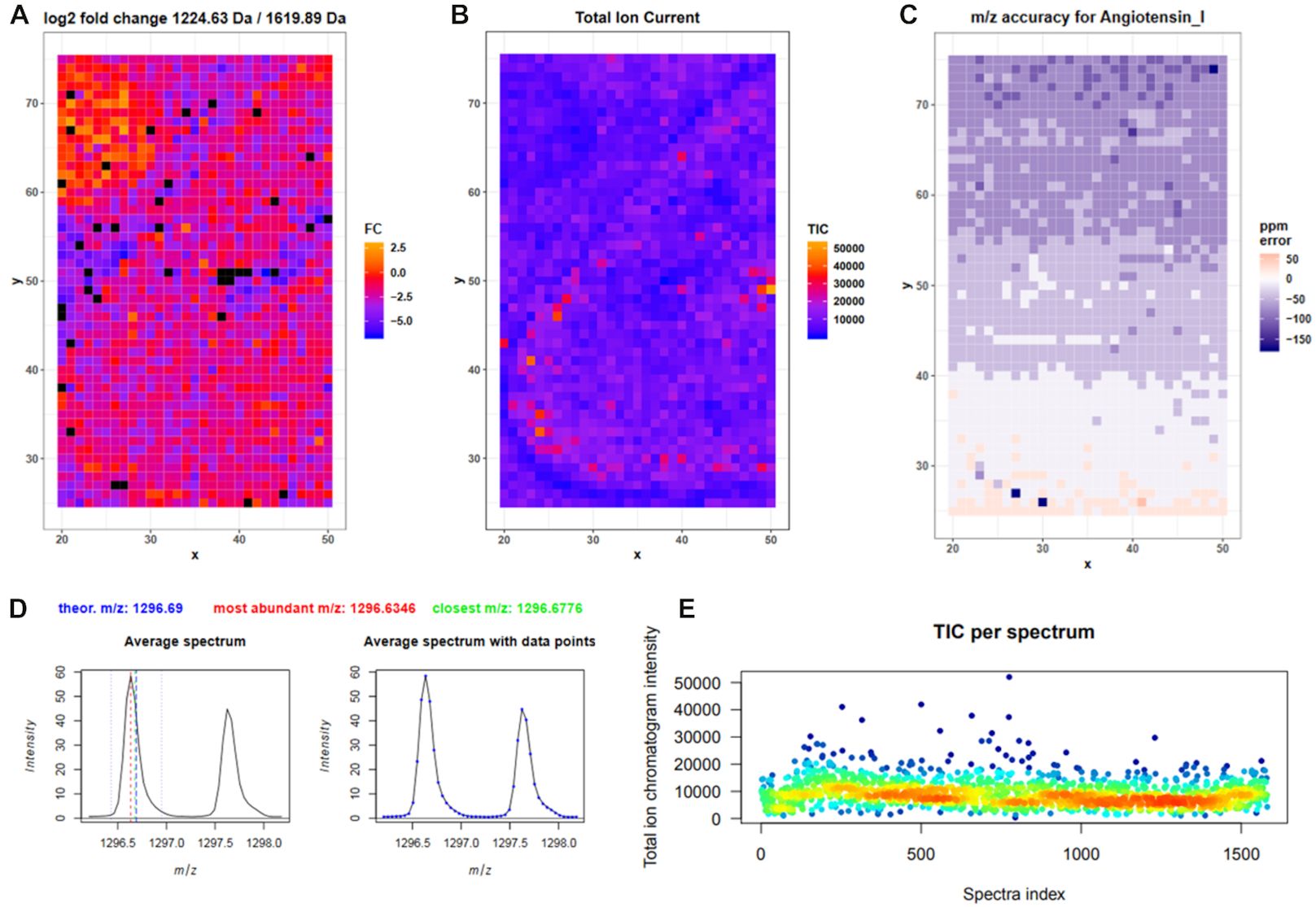
We have integrated 18 dedicated mass spectrometry imaging tools into the Galaxy framework to allow accessible, reproducible, and transparent data analysis. Our tools are based on Cardinal, MALDIquant, and scikit-image and enable all major MSI analysis steps such as quality control, visualization, preprocessing, statistical analysis, and image co-registration. Furthermore, we created hands-on training material for use cases in proteomics and metabolomics. To demonstrate the utility of our tools, we re-analyzed a publicly available N-linked glycan imaging dataset. By providing the entire analysis history online, we highlight how the Galaxy framework fosters transparent and reproducible research.
The Galaxy framework has emerged as a powerful analysis platform for the analysis of MSI data with ease of use and access, together with high levels of reproducibility and transparency.