Cluster Maintenance April 8 and 9
Starting from April 8th 2024, maintenance work will take place at our compute cluster, however our headnode servers are not affected. This means:
Welcome to Galaxy Proteomics
This European Galaxy branches off from the main usegalaxy.eu server and focuses on one specific domain: Proteomics.
Galaxy Proteomics is a multiple-'omics' data-analysis platform with particular emphasis on mass spectrometry-based proteomics. It is hosted by GalaxyEU and maintained by the GalaxyP team from the University of Minnesota, USA.
GalaxyP Resources:
About Us
Publications
Conference presentations
Workshops
Table of Contents
Tutorials
Are you new to Galaxy, or returning after a long time, and looking for help to get started? Take a guided tour through Galaxy's user interface.
Want to learn more about proteomics? Check out the following lesson tutorials from the Galaxy Training Network:
Gateways
Other specialized Galaxy servers curated by the Galaxy-P team containing tools, workflows, and hands-on/video tutorials.
| Server | Tutorial(s) | Video |
|---|---|---|
| Proteogenomics Gateway | Proteogenomics 1: Database creation Proteogenomics 2: Database search Proteogenomics 3: Novel peptide analysis | 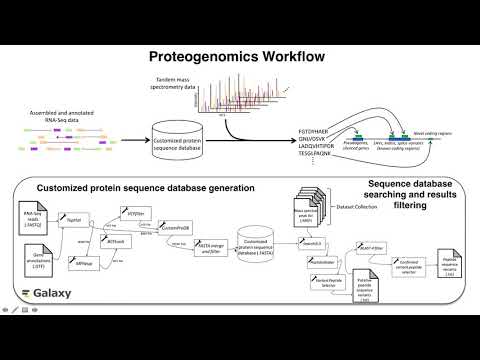 |
| Metaproteomics Gateway | Metaproteomics tutorial |  |
Research Projects
Featured Galaxy-P tools supported with manuscripts, visualization plugins, docker containers, and hands-on tutorials.
| Tool | Description | Paper | Tutorial | Docker |
|---|---|---|---|---|
| CRAVAT-P | Stands for Cancer Related Analysis of Variants Toolkit for Proteogenomics. Submits, checks for, and retrieves data for cancer annotation | Sajulga et. al 2018 | ||
| QuanTP | Correlation between protein and transcript | Kumar et. al 2019 | ||
| MetaQuantome | Quantitative analysis of the function and taxonomy of microbiomes and their interaction |
News and Events
Contributors
Our Data Policy
| Registered Users | Unregistered Users | FTP Data | GDPR Compliance |
|---|---|---|---|
| User data on UseGalaxy.eu (i.e. datasets, histories) will be available as long as they are not deleted by the user. Once marked as deleted the datasets will be permanently removed within 14 days. If the user "purges" the dataset in the Galaxy, it will be removed immediately, permanently. An extended quota can be requested for a limited time period in special cases. | Processed data will only be accessible during one browser session, using a cookie to identify your data. This cookie is not used for any other purposes (e.g. tracking or analytics). If UseGalaxy.eu service is not accessed for 90 days, those datasets will be permanently deleted. | Any user data uploaded to our FTP server should be imported into Galaxy as soon as possible. Data left in FTP folders for more than 3 months, will be deleted. | The Galaxy service complies with the EU General Data Protection Regulation (GDPR). You can read more about this on our Terms and Conditions. |
